OUR TEAM

![]()
![]()
M. Madan Babu, PhD, FRSC
Member, Structural Biology Department
Endowed Chair in Biological Data Science
Director of the Center for Data Driven Discovery
madan.babu@stjude.org
Madan is interested in data science approaches to make biological discoveries and reveal fundamental principles of biological systems. He employs and develops computational and experimental methods to study biological systems at different scales of complexity: from systems at the atomic level (e.g., structures), to the network of interactions between molecules (e.g., protein interaction networks), to the level of an entire population (e.g., variation in the human population and cancer genomes). He envisions that his group’s research findings will help us better understand how mutations cause cancer and other diseases. This knowledge can be used in biotechnology, drug discovery, and personalized medicine.
Before joining St. Jude, Madan was a Programme Leader at the MRC Laboratory of Molecular Biology in Cambridge, UK. His group joined the Center for Data Driven Discovery, and the Structural Biology Department at St. Jude in July 2020. Please see Research and About Madan for more details.

![]()
Angie Williams
Executive Assistant
angie.williams@stjude.org
Angie Williams brings 35 years of experience in administration, event planning, and customer service. She has 13 years of cumulative experience at St. Jude. She started her career as a clerk in 1982 at the Memphis Police Department. Angie joined St. Jude Children’s Research Hospital in 2002 in the Department of Diagnostic Imaging. In her first eight years at St. Jude, she actively supported several doctors, including the Chair of Oncology and the Chair of Chemical Biology and Therapeutics. For the next six years, Angie was Curriculum Manager and Executive Assistant to the Vice President of Academic and Student Services at Colorado Community College. While in Colorado, she fine-tuned her skillset in planning large meetings and symposia.
She returned to St. Jude in 2016 in a dual position in the Cancer Center and the Division of Molecular Oncology under EVP Charles Roberts, MD, Ph.D. She moved to Structural Biology in 2018.

![]()
Ben Leslie, PhD
Director of Lab Operations
ben.leslie@stjude.org
Ben Leslie is a Chemical Biologist whose research include Organic Chemistry, Molecular/ Cell Biology and Biophysics. He is involved in structural and functional studies of GPCR targets that emerge from data driven drug discovery efforts. Additionally, he works on screening projects to discover functional motifs in disordered proteins. Ben earned a PhD in Organic Chemistry with Paul Hergenrother’s group at the University of Illinois where he specialized in Oncology Drug Discovery.
He was a postdoc and later Research Specialist in Taekjip Ha’s Single Molecule Biophysics group.

![]()
![]()
Balaji Santhanam, PhD
Principal Computational Research Scientist
balaji.santhanam@stjude.org
Balaji’s primary research interest is in understanding protein-protein interactions in various key cellular regulatory pathways such as Ubiquitin systems, GPCR pathway, Transcription and Protein kinase signalling process. Specifically, how these interactions are altered in disease states such as cancer, infectious diseases and metabolic disorders and identifying key components of above regulatory pathways for therapeutic intervention strategies.
Balaji obtained his PhD from Molecular Biophysics Unit at Indian Institute of Science, Bangalore, India. Before joining St Jude, Balaji was leading bioinformatics collaborations across the divisions in MRC Laboratory of Molecular Biology, Cambridge, UK.

![]()
Benjamin Lang, PhD
Senior Bioinformatics Research Scientist
benjamin.lang@stjude.org
Benjamin’s research is focussed on identifying features in intrinsically unstructured regions that can control the localisation of proteins within cells, and how these systems might be impaired in cancer and in genetic diseases. Benjamin trained in general and molecular biology in Münster, Zürich and Heidelberg before completing a PhD degree on the evolution of post-translational signalling systems at the MRC-LMB and the University of Cambridge.
Following postdoctoral EIPOD and Marie Skłodowska-Curie IF fellowships at EMBL Heidelberg and at the Centre for Genomic Regulation (CRG) in Barcelona, where he investigated protein linear motifs and protein–RNA interactions, he is currently a Senior Scientist in the group.

![]()
![]()
Cristina Guibao
Scientist
cristina.guibao@stjude.org
Cristina is a Biologist who has worked in the Structural Biology field for most of her career. She is involved in cell-based validation of targets that arise from data-driven discovery projects in the group. Cristina also performs functional validation of GPCR and other targets that emerge from our data mining efforts.
Cristina earned her B.S in biology from Rhodes College.

![]()
Preethi Badrinarayan
Bioinformatics Research Scientist
preethi.badrinarayan@stjude.org
The main focus of Preethi’s research is to study the activation mechanisms, allostery, and dynamics to understand the regulation of kinases in cancer and design target selective kinase drugs. She has joined as part of The Blue-Sky project “Seeing the Invisible in Protein Kinases” at St Jude. She will use inter-disciplinary approaches to characterize the distinct conformational states populated by kinases and to dissect the mechanisms of allosteric regulation and those underpinning oncogenic and drug-resistance mutations.
Before joining St Jude, Dr. Preethi Badrinarayan has received her independent Wellcome-Trust Early Career Grant and has been a principal investigator for the past five years.

![]()
Duccio Malinverni, PhD
Senior Bioinformatics Analyst
duccio.malinverni@stjude.org
Duccio’s research is focused on developing machine learning based approaches to design novel interacting proteins, study protein conformational heterogeneity and investigate the sequence-structure relationship in proteins. In his current work, he focuses on the design of orthogonal protein interactions, with particular interest on membrane receptors. He is also interested in how to use very large biological datasets to study conformational changes in protein kinases.
Duccio obtained his PhD in Physics at the École Polytechnique Fédérale de Lausanne. Before joining St.Jude, he was a post-doctoral researcher at the MRC LMB in Cambridge UK, supported by a Swiss SNF fellowship.
Besian’s research focuses on developing and applying massively parallel in silico screening protocols to rapidly dock billions of commercially available compounds to target proteins in a pipeline designed to accelerate drug discovery and development. He is interested in recognizing druggable protein targets, identifying compound leads with promising therapeutic and pharmacokinetic profiles and move these molecules to proof-of-concept studies in preclinical disease models. He also works in building interactive user-friendly applications to allow for real-time exploratory analysis of scientific data.
Besian obtained his PhD in Biophysical Chemistry at the Centre for Molecular Simulation at the University of Calgary, where he applied multiscale methods to study the reciprocal interplay of membrane proteins with their surrounding lipid environment. He has a master’s degree in pharmacy from the University of Prishtina.

![]()
Xiaohan Li, PhD
Investigator Scientist
xli@mrc-lmb.cam.ac.uk
Xiaohan is currently an investigator scientist supported by a Marie-Curie individual fellowship. He is interested in understanding dynamics and cellular functions of intrinsically disordered proteins and regions using integrative structural, cellular and computational characterisations. His work will provide insight into our understanding of sequence-to-function paradigm for disordered proteins and regions.
Xiaohan got his PhD in biophysical chemistry from Yale University studying how the interaction between intrinsically disordered protein tau and tubulin contributes to tau’s function.

![]()
Maria Marti Solano, PhD
Postdoctoral Fellow
mmarti@mrc-lmb.cam.ac.uk
Maria’s research is focused on understanding how membrane protein structural variation results in functional diversity. In her current work, she employs computational systems biology to explore how membrane protein structural, spatial and interindividual changes can result in differences in physiology and drug response. Maria did her PhD in Biomedicine at the Pompeu Fabra University in Barcelona.
Before joining the group, she was a post-doctoral researcher at the Philipps University Marburg, Germany. She initially joined the MRC LMB supported by a FEBS Long-Term Postdoctoral Fellowship and is currently a Marie Skłodowska-Curie Fellow.

![]()
Assaf Elazar, PhD
Postdoctoral Fellow
assaf.elazar@stjude.org
Assaf’s research project aims to uncover the sequence determinants driving reversible/(i)reversible higher-order assembly of Intrinsic Disordered Regions(IDRs) using an innovative targeted library generation framework, coupled with a high-throughput selection approach.
Assaf obtained his Ph.D. in protein design from the Weizmann Institute Of Science, where he worked on designing membrane proteins. He is currently an EMBO long-term fellow.

![]()
Manbir Sandhu, PhD
Postdoctoral Fellow
manbir.sandhu@stjude.org
Manbir is studying how genetic and post-transcriptional regulation of signaling proteins can illicit functional variation in their activity profiles. This research will shed light on how drug response is modulated in various target tissues due to the heterogeneous nature of gene expression and regulation.
Manbir received his PhD in Computational & Structural Biology from the City of Hope National Cancer Center in Los Angeles, CA before joining the Babu Group at St. Jude Children’s Research Hospital.

![]()
Daniel Estevez Prado, PhD
Postdoctoral Fellow
daniel.estevezprado@stjude.org
Daniel’s project aims to shed light on the biogenesis of polytopic membrane proteins such as G-protein coupled receptors (GPCRs); how the emerging polypeptide inserts into the lipid bilayer, adapts a functional conformation and is eventually expressed at the cell’s surface. He employs an interdisciplinary computational approach investigating the structural, biophysical and genomic signatures associated with GPCR folding, function and misfolding.
Daniel received a Master’s degree in Molecular Biophysics from King’s College London and conducted research as a Postgraduate Research Fellow at the Yale School of Medicine before joining the group.

![]()
Alex Gunnarsson, PhD
Postdoctoral Fellow
alexander.gunnarsson@stjude.org
Alex is interested in how short linear motifs within protein intrinsic disorder allow cells to process complex information and to regulate cellular decision making. The aim of his research is to characterise how the motifs evolve and change over time and how this process enables functional adaptation through motif-mediated network rewiring.
Alex obtained a BA and MSci in Biochemistry from the University of Cambridge working on super resolution image analysis and cryo-EM method development respectively. His work is funded by the MRC LMB.

![]()
Yonathan Goldtzvik, PhD
Postdoctoral Fellow
goldtzvik@mrc-lmb.cam.ac.uk
Yonathan’s research aims to understand the extent of heterogeneity in the cellular machinery that is involved in the production and degradation of proteins. The aim of this project is to understand how variation in sequence and composition of molecular machines involved in these processes affects the regulation of proteins in the cell.
Yonathan obtained his PhD in Biophysics from the University of Maryland College Park where he worked on the biophysics of motor proteins. He is currently an EMBO long-term fellow.

![]()
Greg Słodkowicz, PhD
Postdoctoral Fellow
gslodko@mrc-lmb.cam.ac.uk
Greg is an AstraZeneca-funded postdoc working on elucidating the role of alternative splicing in healthy tissue and in disease, focusing on prostate cancer. Originally trained as a computer scientist, Greg obtained an MSc in Systems Biology while in the group of Chris Workman at the Technical University of Denmark. He then completed a PhD in Molecular Evolution with Nick Goldman at EMBL-EBI, and continues to be interested in evolutionary aspects of the biological systems he studies.
Before joining Madan’s lab, Greg spent one year as a postdoc in the lab of Leo James at the LMB.

![]()
Franziska Marie Heydenreich, PhD
Postdoctoral Fellow
fheyden@stanford.edu

![]()
Sarah Sherman
PhD Student
sarah.sherman@stjude.org
Sarah’s research focus is leveraging experimental and computational approaches for the functional characterization of intrinsically disordered regions (IDRs) within proteins, specifically investigating how these regions can play a role in regulating subcellular localization. She is interested in extending these characterizations to investigate how the mutation of these elements impacts their functioning in diseases such as cancer.
Sarah has a B.S. in Biology and English from Northeastern University. Before joining the group, she worked as a research technician to drive the development of a protease-based diagnostic platform at Glympse Bio. She has also conducted research in Botswana on the diagnostic turn-around times of cervical cancer specimens under a Fulbright grant. In her free time, she enjoys reading and ballroom dance.
Jaimin’s research in the group focuses on developing web tools to visualize and characterize conformational states of proteins from medically important protein families, such as kinases and GPCRs. He also updates and maintains some of the web tools developed by the Babu Group.
Jaimin received a MS from George Mason University in Fairfax, VA, where his thesis project focues on designing a bioinformatics web-tool for miRNA-mRNA interactions analysis in Drosophila. He joined St Jude in 2019 as a Bioinformatics Portal Engineer, before moving to the Babu Group in 2022.

![]()
![]()
Georgi Kanev, PhD
Bioinformatics Research Scientist
georgi.kanev@stjude.org
The main focus of Georgi’s research is building and applying data science and computational approaches that integrate and explore biomolecular structural and bioactivity data. The current focus of his research is to systematically characterize the structural conformational landscape in order to study medically relevant protein families, such as kinases and GPCRs.
Georgi received his PhD from Division of Medicinal Chemistry at Vrije Universiteit Amsterdam, The Netherlands, and Cancer Center Amsterdam, The Netherlands and was a postdoc in the group of Bart Westerman, also at Cancer Center Amsterdam. Prior to joining the Babu Group, Georgi worked as a machine learning scientist for InterAx Biotech AG in Villigen, Switzerland, focusing on development of a specialized virtual screening and machine learning pipeline.

![]()
Hadi Hosseini, PhD
Postdoctoral Research Associate
hadi.hoessini@stjude.org
Hadi holds MS and PhD degrees in Electrical Engineering in the field of control systems with artificial intelligence and was a faculty member at Azad University for 5 years. He also holds an MS in Bioinformatics from the University of Memphis. For the latter, his work focused on correlations between transcriptome data and intraocular diseases such as glaucoma. As a machine learning engineer, he is analyzing the genomics and proteomics data to design prediction models for cancer disease. He also has experience in image processing and computer vision.

![]()
Kalyan Immadisetty, PhD
Bioinformatics Research Scientist
kalyan.immadisetty@stjude.org
Working in conjunction with the Krenciute’s lab in the Department of Bone Marrow Transplant and Cellular Therapy, Kalyan’s research focuses on leveraging the state-of-the-art computational tools including molecular dynamics (MD) simulations, and machine learning to design functional CAR T-cell receptors for treating cancer. He received a PhD from Duquesne University in Pittsburgh, followed by a postdoc at the University of Arkansas Fayetteville. Prior to joining St Jude, Kalyan was a Research Scientist at the Stritch School of Medicine and Loyola University in Chicago, where he used MD simulations, free energy calculations and machine learning to study various therapeutically important protein targets.

![]()
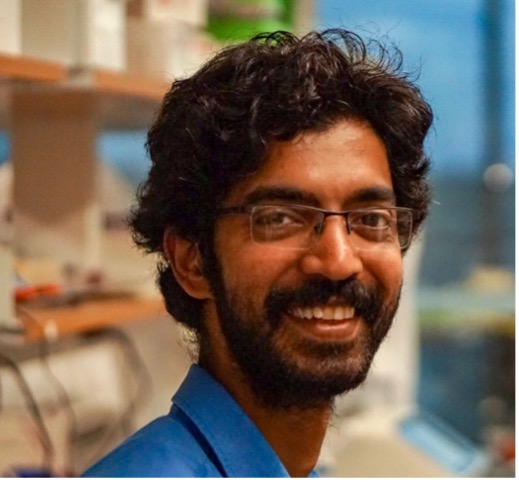
Vikas Trivedi, PhD
Scientist
vikas.trivedi@stjude.org
Vikas’ research focuses on the cellular validation of targets indicated by data science methods, designing and conducting experiments to support findings. He is an expert in synthetic biology, functional genomics, developing biosensors, highthroughput protein engineering using deep mutational scanning, enzymology, metabolic engineering in yeast.
Dr Trivedi earned his PhD at the Indian Institute of Technology – Bombay, investigating microbial degradation of xenobiotics at the biochemical and molecular level. He then completed a postdoctoral fellowship at Tufts University, working on projects including developing peptide probes for microbiota, yeast metabolic engineering for non-native substrate utilization, and enzyme engineering for phenylketonuria.

![]()
Vyoma Sheth, MS
Senior Bioinformatics Analyst
vyoma.sheth@stjude.org
Vyoma’s research is focused on understanding the transcriptional and post- transcriptional regulation in cancer. Also, develop the drugs that focused on targeted degradation of undruggable oncoprotein.
Vyoma received her MS in Bioinfromatics and Computational Biology from the University of South Florida – Morsani College of Medicine, where her work focused on the epithelial – mesenchymal transition in cancer She joined the Babu Group from the Channing Division at Brigham and Women’s Hospital in Boston.

![]()
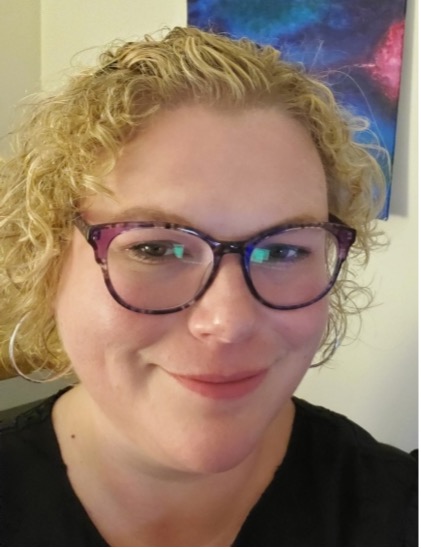
Elly Park, PhD
Director, Administration and Operations
elly.park@stjude.org
Elly holds a PhD in Biochemistry and Molecular Biology, during which time she studied molecular mechanisms of prostate cancer metastasis. She joins St Jude in 2022 following ~7years as an Associate Professor at Southwest Tennessee Community College in Memphis.

![]()
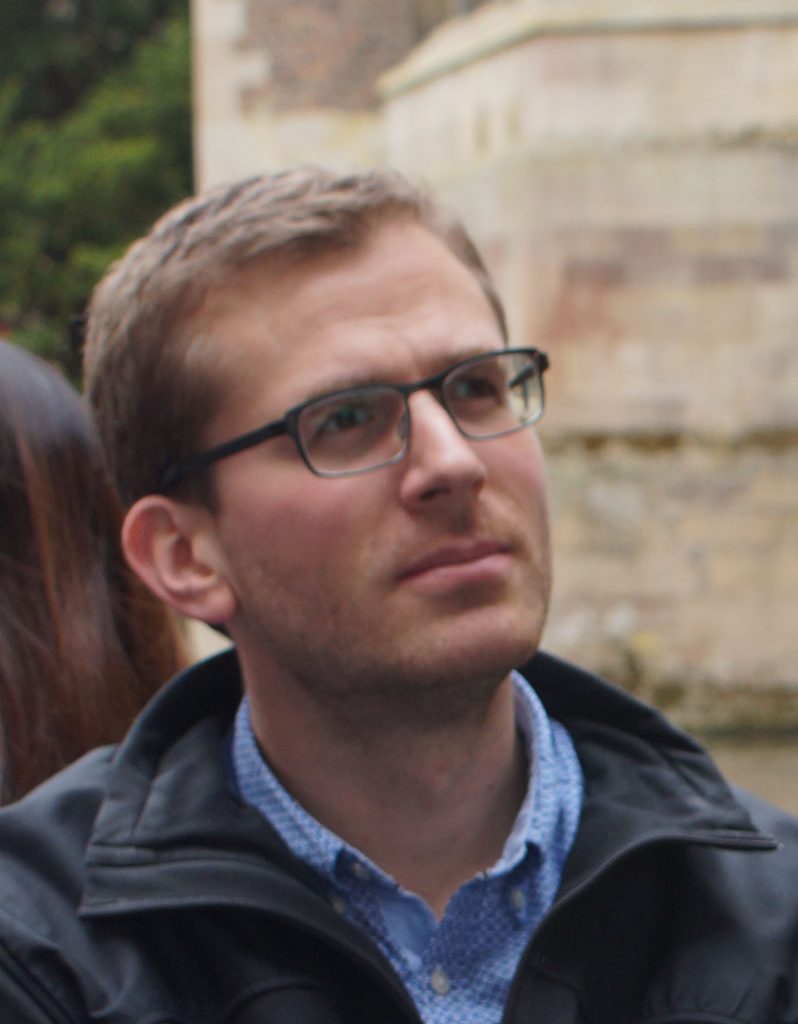
Hannes Harbrecht
PhD student
hannesh@mrc-lmb.cam.ac.uk

![]()
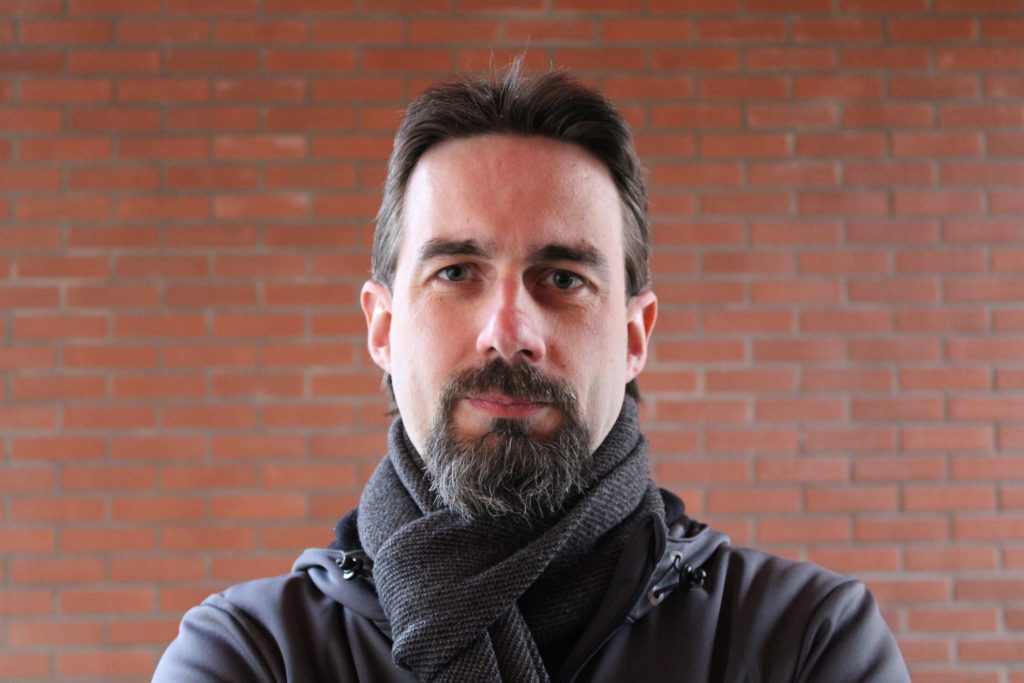
Bálint Mészáros, PhD
Principal Computational Research Scientist
balint.meszaros@stjude.org
Balint Bio